Filter nodes using dplyr-style expressions on any node column or centrality
measure. Returns a cograph_network object by default (universal format), or
optionally the same format as input when keep_format = TRUE.
Arguments
- x
Network input: cograph_network, matrix, igraph, network, or tna object.
- ...
Filter expressions using any node column or centrality measure. Available variables include:
- Node columns
All columns in the nodes dataframe:
id,label,name,x,y,inits,color, plus any custom- Centrality measures
degree,indegree,outdegree,strength,instrength,outstrength,betweenness,closeness,eigenvector,pagerank,hub,authority
Examples:
degree >= 3,label %in% c("A", "B"),pagerank > 0.1 & degree >= 2.- .keep_edges
How to handle edges. One of:
"internal"(default) Keep only edges between remaining nodes
"none"Remove all edges
- keep_format
Logical. If TRUE, return the same format as input (matrix returns matrix, igraph returns igraph, etc.). Default FALSE returns cograph_network (universal format).
- directed
Logical or NULL. If NULL (default), auto-detect from matrix symmetry. Set TRUE to force directed, FALSE to force undirected. Only used for non-cograph_network inputs.
Value
A cograph_network object with filtered nodes. If keep_format = TRUE,
returns the same type as input (matrix, igraph, network, etc.).
See also
filter_edges, splot, subset_nodes
Examples
adj <- matrix(c(0, .5, .8, 0,
.5, 0, .3, .6,
.8, .3, 0, .4,
0, .6, .4, 0), 4, 4, byrow = TRUE)
rownames(adj) <- colnames(adj) <- c("A", "B", "C", "D")
# Keep only high-degree nodes (returns cograph_network)
filter_nodes(adj, degree >= 3)
#> Cograph network: 2 nodes, 1 edges ( undirected )
#> Source: filtered
#> Nodes (2): B, C
#> Edges: 1 / 1 (density: 100.0%)
#> Weights: [0.300, 0.300] | mean: 0.300
#> Strongest edges:
#> B -- C 0.300
#> Layout: none
# Keep format: matrix in, matrix out
filter_nodes(adj, degree >= 3, keep_format = TRUE)
#> B C
#> B 0 0.3
#> C 0 0.0
# Filter by node label
splot(filter_nodes(adj, label %in% c("A", "C")))
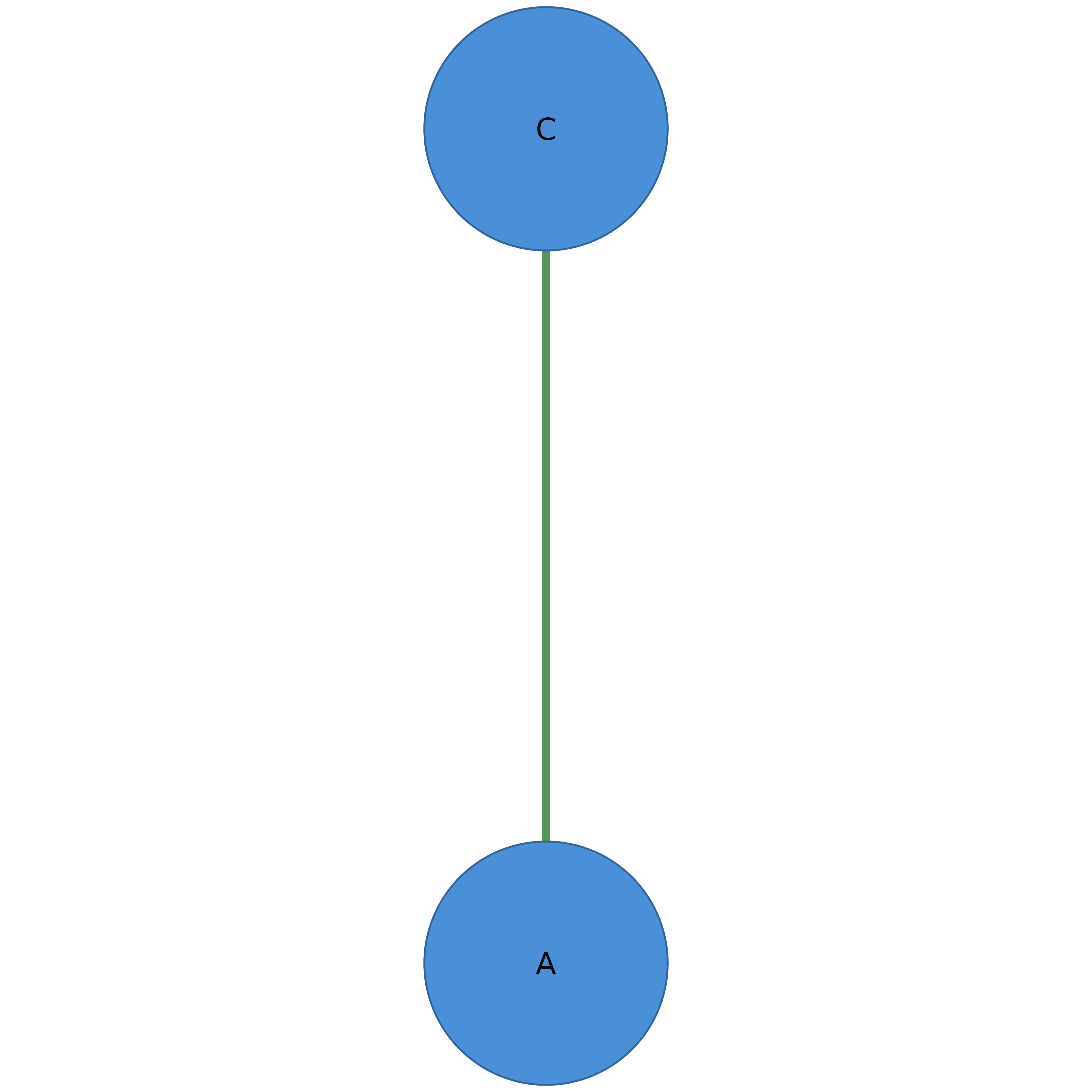 # Combine centrality and metadata filters
splot(filter_nodes(adj, degree >= 2 & label != "D"))
# Combine centrality and metadata filters
splot(filter_nodes(adj, degree >= 2 & label != "D"))
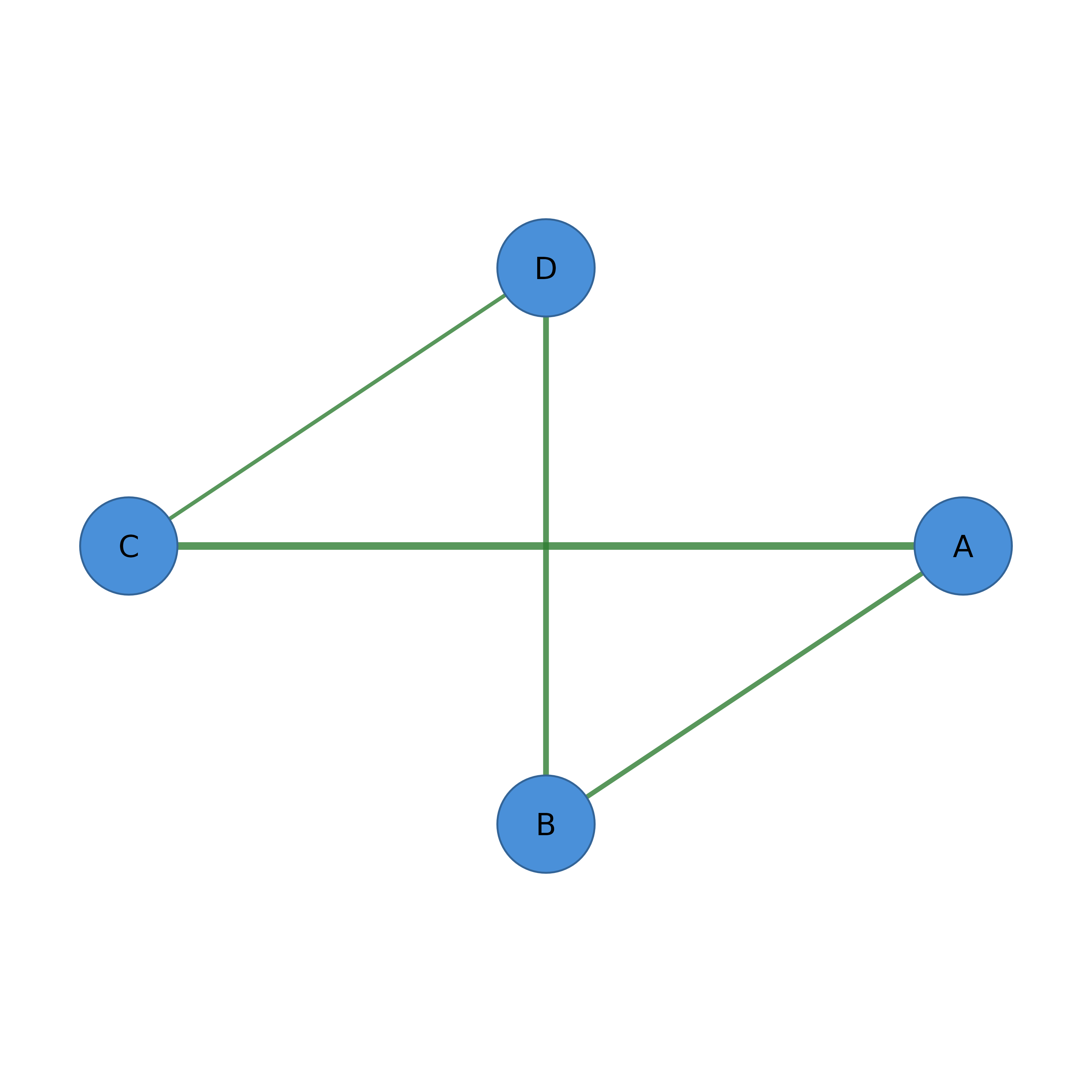
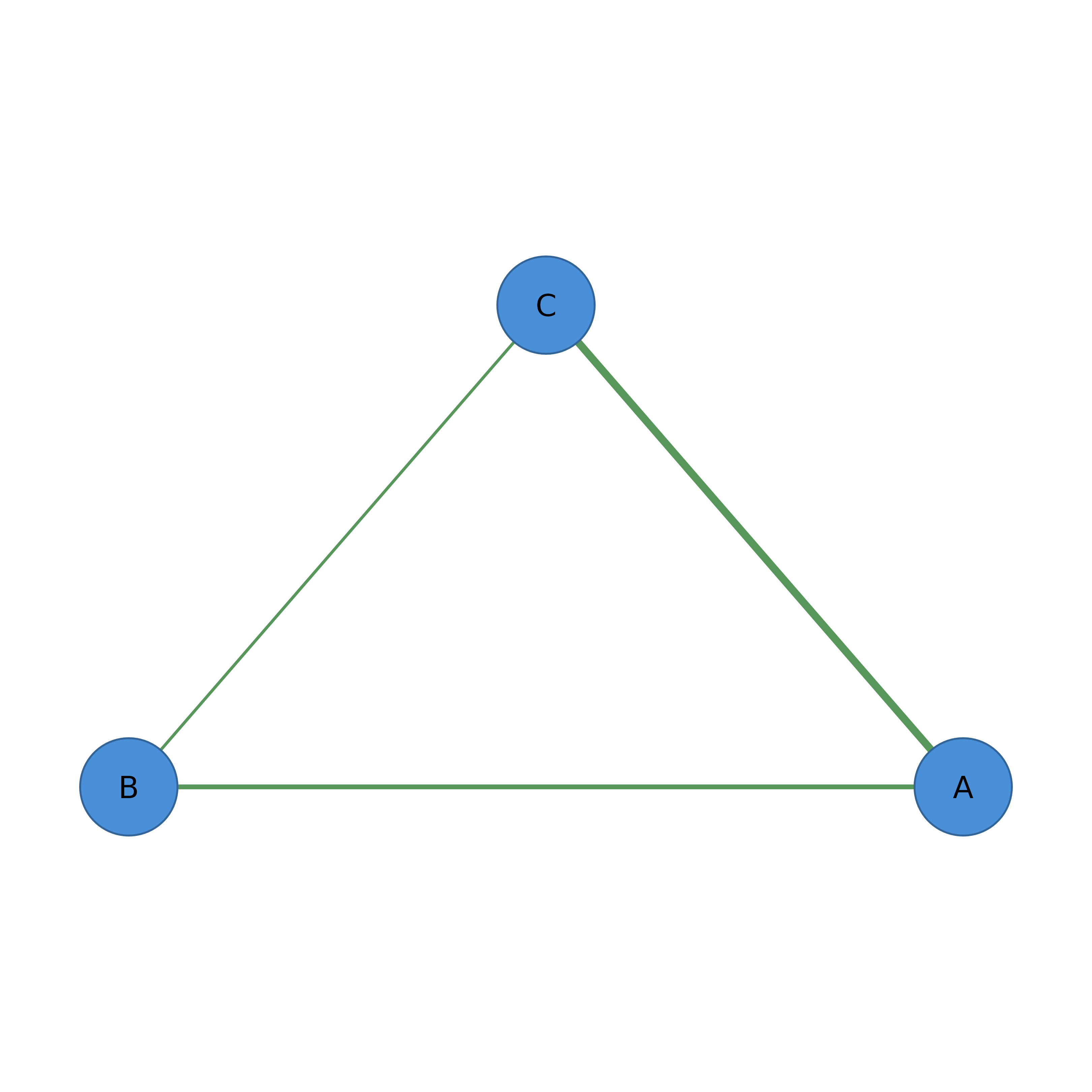 # With cograph_network (pipe-friendly)
net <- as_cograph(adj)
net |>
filter_edges(weight > 0.3) |>
filter_nodes(degree >= 2) |>
splot()
# With cograph_network (pipe-friendly)
net <- as_cograph(adj)
net |>
filter_edges(weight > 0.3) |>
filter_nodes(degree >= 2) |>
splot()
 # With igraph (keep_format = TRUE returns igraph)
if (requireNamespace("igraph", quietly = TRUE)) {
g <- igraph::make_ring(5)
filter_nodes(g, degree >= 2, keep_format = TRUE) # Returns igraph
}
#> IGRAPH 34a80ac UNW- 5 5 --
#> + attr: name (v/c), weight (e/n)
#> + edges from 34a80ac (vertex names):
#> [1] 1--2 2--3 3--4 4--5 1--5
# With igraph (keep_format = TRUE returns igraph)
if (requireNamespace("igraph", quietly = TRUE)) {
g <- igraph::make_ring(5)
filter_nodes(g, degree >= 2, keep_format = TRUE) # Returns igraph
}
#> IGRAPH 34a80ac UNW- 5 5 --
#> + attr: name (v/c), weight (e/n)
#> + edges from 34a80ac (vertex names):
#> [1] 1--2 2--3 3--4 4--5 1--5
