Do we need another network package?
R already has igraph for computation and qgraph for psychometric networks. tidygraph tries to unify things with a dplyr grammar. So why write another one?
The short answer: none of these packages let you go from a raw matrix to a filtered, annotated, publication-ready figure without switching between ecosystems, reformatting objects, or writing boilerplate. cograph tries to close that gap.
This document walks through the specific things cograph does differently, with working examples.
Filtering and selecting — like data frames
In igraph, subsetting a network means calling
induced_subgraph(), delete_edges(), or
indexing into V(g) and E(g). In tidygraph, you
need activate(nodes) |> filter(...). Both approaches
require you to think about the object structure.
cograph lets you filter nodes and edges with expressions that look
like subset():
mat <- matrix(c(
0.0, 0.5, 0.8, 0.1, 0.0,
0.3, 0.0, 0.2, 0.6, 0.4,
0.7, 0.1, 0.0, 0.3, 0.5,
0.0, 0.4, 0.2, 0.0, 0.9,
0.1, 0.3, 0.6, 0.8, 0.0
), 5, 5, byrow = TRUE)
rownames(mat) <- colnames(mat) <- c("Read", "Write", "Plan", "Code", "Test")
net <- as_cograph(mat)Keep only edges above a threshold:
strong <- filter_edges(net, weight > 0.5)
get_edges(strong)
#> from to weight
#> 1 3 1 0.7
#> 2 1 3 0.8
#> 3 5 3 0.6
#> 4 2 4 0.6
#> 5 5 4 0.8
#> 6 4 5 0.9Keep nodes that have high degree and high PageRank — the centrality measures are computed on the fly:
hubs <- filter_nodes(net, degree >= 3 & pagerank > 0.15)
get_nodes(hubs)
#> id label name x y
#> 1 1 Read Read NA NA
#> 2 2 Write Write NA NA
#> 3 3 Plan Plan NA NA
#> 4 4 Code Code NA NA
#> 5 5 Test Test NA NAStructural selections
select_nodes() goes further. It knows about components,
neighborhoods, articulation points, and k-cores — and it computes them
lazily (only what your expression actually references):
# Top 3 nodes by betweenness
top3 <- select_top(net, n = 3, by = "betweenness")
get_nodes(top3)
#> id label name x y
#> 1 1 Write Write NA NA
#> 2 2 Plan Plan NA NA
#> 3 3 Code Code NA NA
# Ego network: everything within 1 hop of "Code"
ego <- select_neighbors(net, of = "Code", order = 1)
get_nodes(ego)
#> id label name x y
#> 1 1 Read Read NA NA
#> 2 2 Write Write NA NA
#> 3 3 Plan Plan NA NA
#> 4 4 Code Code NA NA
#> 5 5 Test Test NA NAFor edges, you can select by structure too:
# Edges involving "Code"
code_edges <- select_edges_involving(net, nodes = "Code")
get_edges(code_edges)
#> from to weight
#> 1 4 2 0.4
#> 2 4 3 0.2
#> 3 1 4 0.1
#> 4 2 4 0.6
#> 5 3 4 0.3
#> 6 5 4 0.8
#> 7 4 5 0.9
# Top 5 edges by weight
top5 <- select_top_edges(net, n = 5)
get_edges(top5)
#> from to weight
#> 1 2 1 0.7
#> 2 1 2 0.8
#> 3 4 2 0.6
#> 4 4 3 0.8
#> 5 3 4 0.9
# Edges between two node sets
between <- select_edges_between(net,
set1 = c("Read", "Write"),
set2 = c("Code", "Test")
)
get_edges(between)
#> from to weight
#> 1 4 1 0.1
#> 2 3 2 0.4
#> 3 4 2 0.3
#> 4 1 3 0.1
#> 5 2 3 0.6
#> 6 2 4 0.4Format in, same format out
If you pass a matrix, you can get a matrix back:
filter_edges(mat, weight > 0.5, keep_format = TRUE)
#> Read Write Plan Code Test
#> Read 0.0 0 0.8 0.0 0.0
#> Write 0.0 0 0.0 0.6 0.0
#> Plan 0.7 0 0.0 0.0 0.0
#> Code 0.0 0 0.0 0.0 0.9
#> Test 0.0 0 0.6 0.8 0.0Same for igraph objects. No lock-in.
Centrality — one call, tidy table
igraph requires a separate function for each centrality measure, and
each returns a different format. page_rank(g) gives you a
list with $vector. betweenness(g) gives a
named numeric. Building a comparison table takes 10+ lines.
In cograph, one call with no arguments returns all 34 measures as a tidy data frame:
centrality(net, digits = 3)
#> node degree_all strength_all closeness_all eccentricity_all coreness_all
#> 1 Read 6 2.5 1.25 0.3 6
#> 2 Write 8 2.8 1.00 0.3 6
#> 3 Plan 8 3.4 1.00 0.4 6
#> 4 Code 7 3.3 1.25 0.3 6
#> 5 Test 7 3.6 1.00 0.4 6
#> harmonic_all diffusion_all leverage_all kreach_all alpha_all power_all
#> 1 26.667 36 -0.110 4 -0.835 -0.885
#> 2 20.000 36 0.069 4 -1.195 -1.046
#> 3 20.833 36 0.069 4 -1.772 -1.046
#> 4 23.333 36 -0.014 4 -2.225 -0.966
#> 5 20.833 36 -0.014 4 -2.366 -1.046
#> betweenness eigenvector pagerank authority hub constraint transitivity
#> 1 3.0 0.582 0.151 0.540 0.736 0.761 0.400
#> 2 3.5 0.674 0.172 0.666 0.800 0.773 0.214
#> 3 5.0 0.882 0.216 0.956 0.725 0.658 0.214
#> 4 4.0 0.966 0.222 1.000 0.763 0.761 0.286
#> 5 0.0 1.000 0.240 0.872 1.000 0.711 0.286
#> subgraph laplacian load current_flow_closeness current_flow_betweenness
#> 1 212.537 88 12.0 0.765 0.221
#> 2 329.433 128 12.5 0.861 0.270
#> 3 329.433 128 14.0 0.917 0.312
#> 4 274.892 102 13.0 0.898 0.201
#> 5 274.892 114 9.0 0.960 0.313
#> voterank percolation
#> 1 0.4 0.250
#> 2 1.0 0.292
#> 3 0.8 0.417
#> 4 0.6 0.333
#> 5 0.2 0.000That includes degree, strength, betweenness, closeness, PageRank, eigenvector, but also less common ones like load, current-flow betweenness, voterank, percolation, diffusion, and leverage — all computed natively, without extra packages.
If you only need a subset:
centrality(net, measures = c("degree", "betweenness", "pagerank"), digits = 3)
#> node degree_all betweenness pagerank
#> 1 Read 6 3.0 0.151
#> 2 Write 8 3.5 0.172
#> 3 Plan 8 5.0 0.216
#> 4 Code 7 4.0 0.222
#> 5 Test 7 0.0 0.240Need just one measure as a named vector? Use the wrapper:
centrality_pagerank(net)
#> Read Write Plan Code Test
#> 0.1507079 0.1715631 0.2157373 0.2223834 0.2396084
centrality_betweenness(net)
#> Read Write Plan Code Test
#> 3.0 3.5 5.0 4.0 0.0You can normalize, sort, and round in the same call:
centrality(net, measures = c("degree", "betweenness", "pagerank"),
normalized = TRUE, sort_by = "pagerank", digits = 3)
#> node degree_all betweenness pagerank
#> 1 Test 0.875 0.0 1.000
#> 2 Code 0.875 0.8 0.928
#> 3 Plan 1.000 1.0 0.900
#> 4 Write 1.000 0.7 0.716
#> 5 Read 0.750 0.6 0.629Edge-level centrality
edge_centrality(net, sort_by = "betweenness", digits = 3)
#> from to weight betweenness
#> 1 Code Plan 0.2 8.0
#> 2 Read Code 0.1 7.0
#> 3 Plan Write 0.1 6.5
#> 4 Write Read 0.3 4.0
#> 5 Test Read 0.1 3.0
#> 6 Write Test 0.4 2.5
#> 7 Plan Test 0.5 1.5
#> 8 Test Write 0.3 1.0
#> 9 Write Plan 0.2 1.0
#> 10 Plan Code 0.3 1.0
#> 11 Plan Read 0.7 0.0
#> 12 Read Write 0.5 0.0
#> 13 Code Write 0.4 0.0
#> 14 Read Plan 0.8 0.0
#> 15 Test Plan 0.6 0.0
#> 16 Write Code 0.6 0.0
#> 17 Test Code 0.8 0.0
#> 18 Code Test 0.9 0.0Network-level summary
One-row data frame with density, diameter, transitivity, centralization, reciprocity, and more:
network_summary(net, digits = 3)
#> node_count edge_count density component_count diameter mean_distance min_cut
#> 1 5 18 0.9 1 0.8 0.35 3
#> centralization_degree centralization_in_degree centralization_out_degree
#> 1 0.125 0.1 0.1
#> centralization_betweenness centralization_closeness centralization_eigen
#> 1 0.028 0 0.097
#> transitivity reciprocity assortativity_degree hub_score authority_score
#> 1 1.009 0.8 -0.25 NA NAAdd detailed = TRUE for mean/sd of node-level measures,
or extended = TRUE for girth, radius, global efficiency,
etc.
Community detection — one call
igraph has cluster_louvain(),
cluster_walktrap(), etc. — different function per
algorithm, inconsistent parameter names. cograph wraps all of them
behind one function with a default:
# Undirected network for community detection
sym <- (mat + t(mat)) / 2
diag(sym) <- 0
cograph::communities(sym)
#> Community structure (louvain)
#> Number of communities: 2
#> Modularity: 0.0985
#> Community sizes: 2, 3
#> Nodes: 5Pick a different algorithm by name, or use two-letter shorthands:
cograph::communities(sym, method = "walktrap")
#> Community structure (walktrap)
#> Number of communities: 2
#> Modularity: 0.0985
#> Community sizes: 2, 3
#> Nodes: 5
com_fg(sym) # fast greedy
#> Community structure (fast_greedy)
#> Number of communities: 2
#> Modularity: 0.0985
#> Community sizes: 3, 2
#> Nodes: 5
com_im(mat) # infomap (works on directed too)
#> Community structure (infomap)
#> Number of communities: 1
#> Modularity: 0
#> Community sizes: 5
#> Nodes: 5If you just want a node-to-community data frame:
detect_communities(sym, method = "walktrap")
#> node community
#> 1 Read 1
#> 2 Write 2
#> 3 Plan 1
#> 4 Code 2
#> 5 Test 2Quality and significance
How good is the partition?
comm <- com_wt(mat)
det <- detect_communities(mat, method = "walktrap")
cluster_list <- split(det$node, det$community)
cqual(mat, cluster_list)
#> Cluster Quality Metrics
#> =======================
#>
#> Global metrics:
#> Modularity: 0
#> Coverage: 1
#> Clusters: 1
#>
#> Per-cluster metrics:
#> cluster cluster_name n_nodes internal_edges cut_edges internal_density
#> 1 1 5 7.8 0 0.39
#> avg_internal_degree expansion cut_ratio conductance
#> 3.12 0 NA 0Is the modularity significantly higher than chance? Permutation test against a null model:
csig(mat, comm, n_random = 200, seed = 1)
#> Cluster Significance Test
#> =========================
#>
#> Null model: configuration (n = 200 )
#> Observed modularity: 0
#> Null mean: 0.1486
#> Null SD: 0.0781
#> Z-score: -1.9
#> P-value: 0.97139
#>
#> Conclusion: No significant community structure (p >= 0.05)Compare two solutions
comm2 <- com_fg(mat)
compare_communities(comm, comm2, method = "nmi")
#> [1] 0Consensus clustering
Run a stochastic algorithm many times and threshold the co-occurrence matrix:
com_consensus(mat, method = "infomap", n_runs = 50, seed = 1)
#> Community structure (consensus_infomap)
#> Number of communities: 1
#> Modularity: 0
#> Community sizes: 5
#> Nodes: 5Format interoperability
cograph accepts matrices, edge lists, igraph, statnet, qgraph, and tna objects natively. And it converts back:
net <- as_cograph(mat)
# To igraph
g <- to_igraph(net)
class(g)
#> [1] "igraph"
# To matrix
m <- to_matrix(net)
m[1:3, 1:3]
#> Read Write Plan
#> Read 0.0 0.5 0.8
#> Write 0.3 0.0 0.2
#> Plan 0.7 0.1 0.0
# To edge list
head(to_df(net))
#> from to weight
#> 1 Write Read 0.3
#> 2 Plan Read 0.7
#> 3 Test Read 0.1
#> 4 Read Write 0.5
#> 5 Plan Write 0.1
#> 6 Code Write 0.4No format lock-in. Use cograph for what it’s good at, convert back when you need something else.
Robustness, motifs, backbone
A few more things that would otherwise require separate packages:
Robustness analysis
How does the network hold up when you remove nodes by betweenness (targeted attack) vs. random failure?
rob <- robustness(mat, measure = "betweenness", strategy = "sequential", seed = 1)
rob_rand <- robustness(mat, measure = "random", n_iter = 50, seed = 1)
# Area under curve — higher means more robust
robustness_auc(rob)
#> [1] 0.5
robustness_auc(rob_rand)
#> [1] 0.5Disparity filter (backbone extraction)
Keep only edges that carry a disproportionate share of a node’s weight:
backbone <- disparity_filter(mat, level = 0.5)
backbone
#> Read Write Plan Code Test
#> Read 0 1 1 0 0
#> Write 0 0 0 1 1
#> Plan 1 0 0 0 1
#> Code 0 1 0 0 1
#> Test 0 1 1 1 0Motif census
Count triad types with significance testing against a configuration model:
motif_census(mat, n_random = 100)
#> Network Motif Analysis
#> Size: 3-node motifs (directed)
#> Null model: configuration (n=100)
#>
#> Significant motifs:
#> motif count expected z p
#> 210 4 1.4 2.02 0.044
#> 300 5 0.1 12.57 <2e-16
#>
#> Over-represented: 2 | Under-represented: 0Working with probabilistic networks
cograph was built with transition networks in mind — matrices where rows sum to 1 and edges are probabilities, not just weights. This matters in a few places.
Smart weight inversion
Path-based centrality measures (betweenness, closeness, harmonic) need distances, not probabilities. For transition networks, a high-probability edge should mean short distance. cograph handles this automatically:
- For tna objects: weights are inverted (distance = 1/probability)
- For other networks: weights are used as-is (standard igraph convention)
No manual toggle needed.
First-class tna support
If you use the tna package for Transition Network Analysis, cograph understands its objects directly:
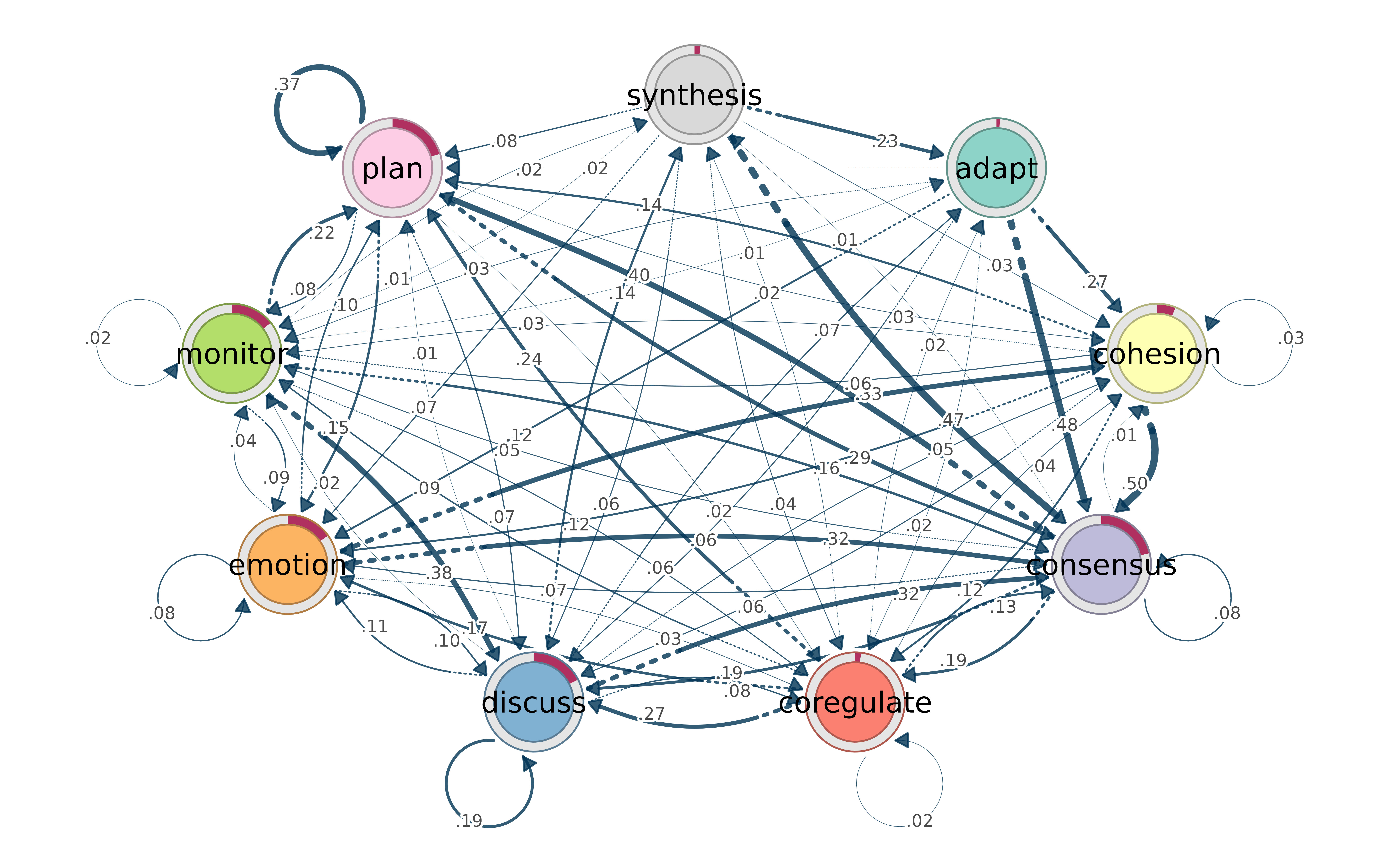
Bootstrap results render automatically — significant transitions as solid edges, non-significant as dashed:
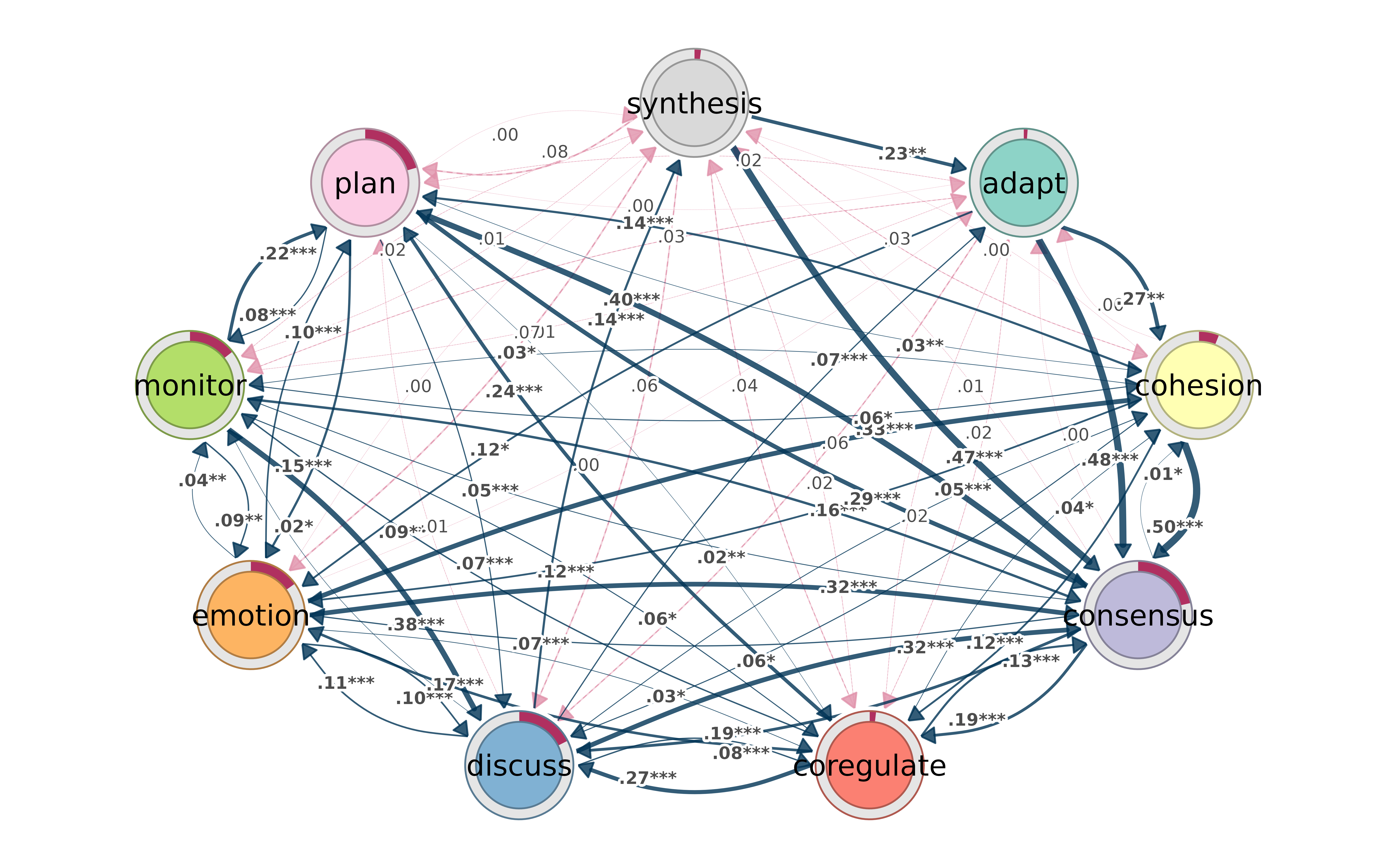
Permutation test results get color-coded by group effect:
model1 <- tna(group_regulation[1:1000,])
model2 <- tna(group_regulation[1001:2000,])
perm <- permutation_test(model1, model2, iter = 1000)
splot(perm)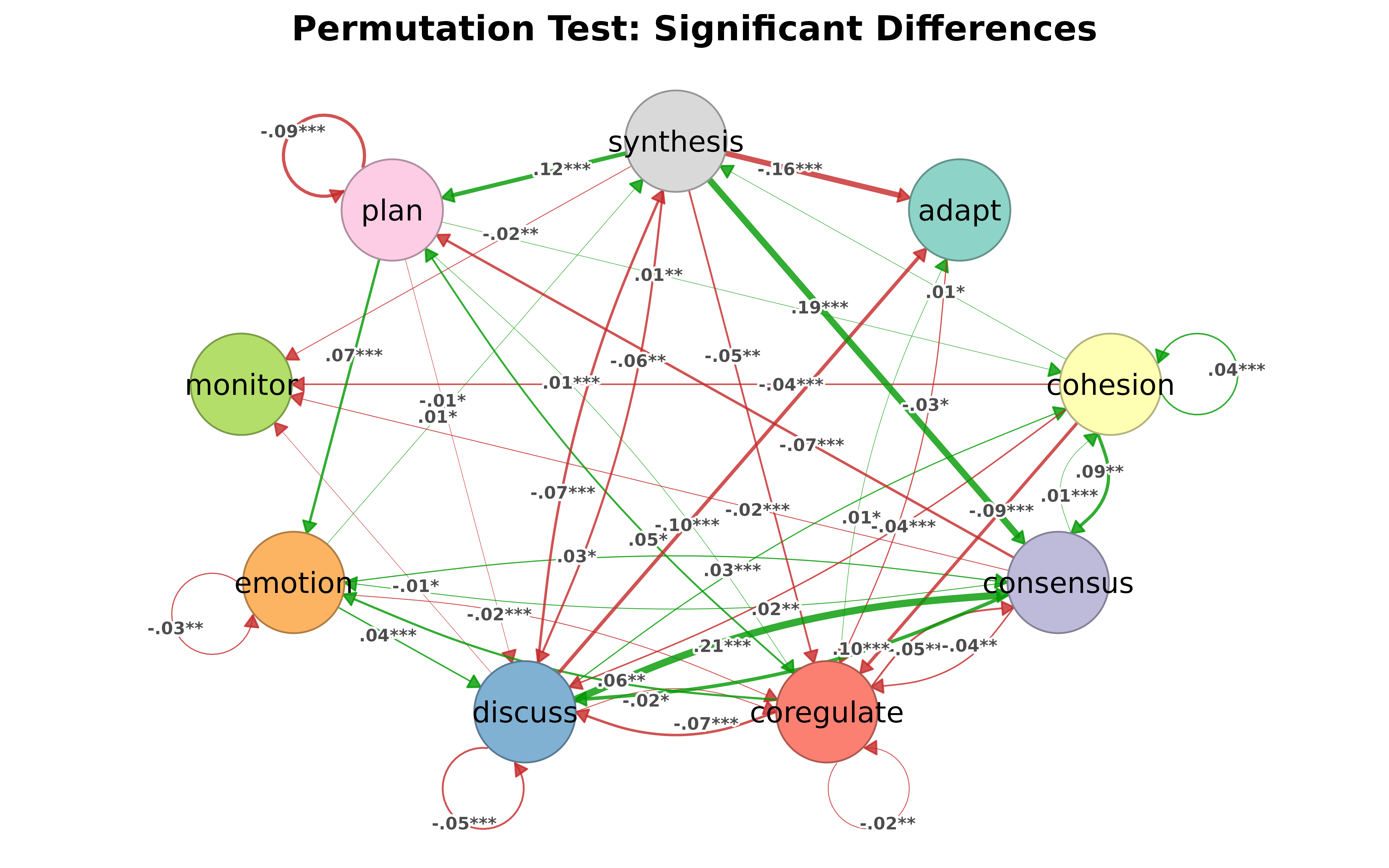
Group comparisons with plot_compare() show element-wise
probability differences, with donuts on nodes for initial state
shifts.
qgraph compatibility
Researchers coming from qgraph can use familiar parameter names —
vsize, asize, edge.color, etc. —
they are translated automatically. When both the cograph name and the
qgraph alias are present, the cograph name wins.
Nestimate integration
cograph also plots Nestimate objects (bootstrap forests, permutation results, glasso networks) without importing the package — dispatch is by class name only.
Summary
cograph is not trying to replace igraph’s graph algorithms or tidygraph’s data manipulation. It fills a different gap: going from data to a filtered, annotated, publication-ready network figure with minimal code, while staying interoperable with everything else in the R network ecosystem.
The main ideas:
- Filter and select with data-frame-like expressions, not object-specific APIs
- Centrality as a tidy data frame, not a list of separate calls
- Community detection behind one function with shorthands, quality metrics, and significance tests
- Statistical annotation (CIs, p-values, stars) directly on the figure
- No format lock-in — matrix/igraph/statnet/tna in and out
- Probabilistic networks handled correctly out of the box
